Creates a split of the row ids of a Task into a training and a test set, and optionally a validation set.
Examples
# regression task partitioned into training and test set
task = tsk("california_housing")
split = partition(task, ratio = 0.5)
data = data.frame(
y = c(task$truth(split$train), task$truth(split$test)),
split = rep(c("train", "predict"), lengths(split[c("train", "test")]))
)
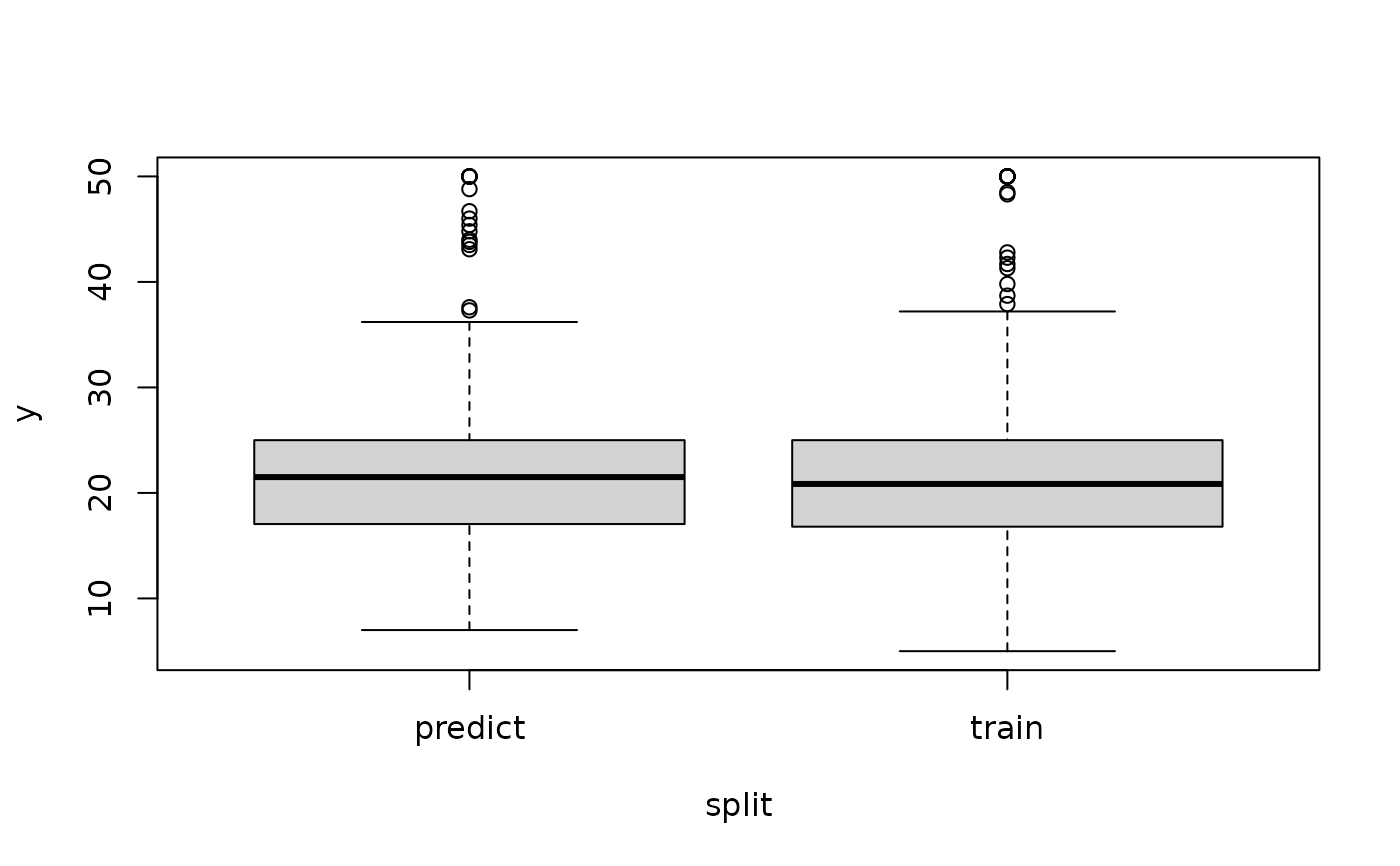
boxplot(y ~ split, data = data)
 # classification task partitioned into training, test and validation set
task = tsk("pima")
split = partition(task, c(0.66, 0.14))
# classification task partitioned into training, test and validation set
task = tsk("pima")
split = partition(task, c(0.66, 0.14))